Lung cancer is the leading cause of cancer worldwide.The major two forms of lung cancer are non small cell lung cancer (NSCLC=85%) of lung cancer and small-cell lung cancer (SCLC=15%). Despite recent advances in the management of resected lung cancer tumors and more effective treatments in the metastatic setting, the cure rate of lung cancer remains low. Successful molecular testing of lung cancer requires the identification and understanding of events that takes place during the multi step tumorigenic process of lung cancer. As with other solid tumors, lung cancer is the result of the accumulation of genetic and epigenetic alterations over a long course of exposure to a carcinogen.
Discovering new prognostic or predictive biomarkers or developing new detection tools for lung cancer is one of the major areas of transitional cancer research. However, given our current understanding of the multifactorial process of lung carcinogenesis and hetrogeneous nature of the disease, monitoring of one or a few genes is limited. A pangenomic analysis seems more efficient for deciphering the complexity of lung cancer. The prospect of identifying specific events in lung carcinogenesis is significantly brightened by the recent development of high-throughput gene expression analysis.
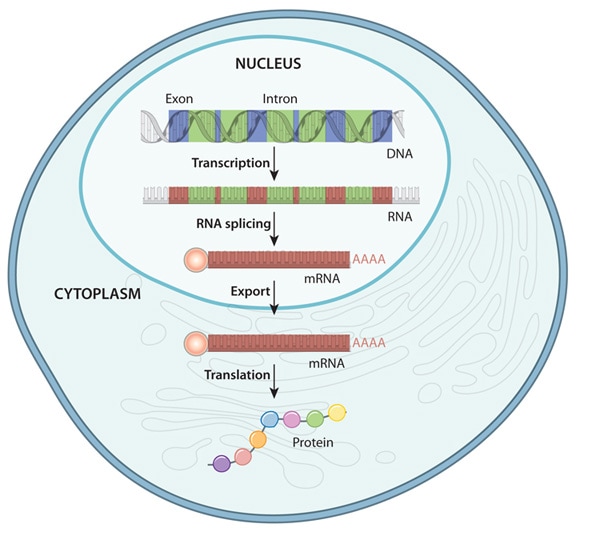
Sherif El-Refai further tried to understand Gene expression, a biological process at the cellular level involves the conversion of DNA information into molecules such as proteins. The process that allowed the cells to adjust to their surroundings by controlling when and how many proteins are to be produced. The gene expression includes two stages: transcription and translation. During transcription, DNA information is copied to produce messenger RNA, which carries the information to protein-producing ribosomes in the cell. Carrier molecules read this information during the translation stage, and ribosomes use it to form a new protein.


 RSS Feed
RSS Feed
